The UMD-THAP1 mutations database
Structural domains and key residues
This function displays the distribution of small rearrangements in structural domains and key residues of the THAP1 protein. Only mutations/variations found in probands are taken into account.
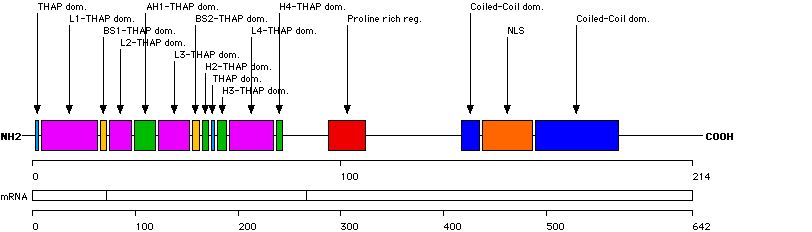
| Structural domain | Number of records |
|---|
| THAP dom. [1-2] | 5 (2.7 %) |
| L1-THAP dom. [3-21] | 28 (14.9 %) |
| BS1-THAP dom. [22-24] | 5 (2.7 %) |
| L2-THAP dom. [25-32] | 19 (10.1 %) |
| AH1-THAP dom. [33-40] | 3 (1.6 %) |
| L3-THAP dom. [41-51] | 29 (15.4 %) |
| BS2-THAP dom. [52-54] | 3 (1.6 %) |
| H2-THAP dom. [55-57] | 5 (2.7 %) |
| THAP dom. [58-59] | 3 (1.6 %) |
| H3-THAP dom. [60-63] | 0 (0 %) |
| L4-THAP dom. [64-78] | 14 (7.4 %) |
| H4-THAP dom. [79-81] | 10 (5.3 %) |
| Proline rich reg. [96-108] | 9 (4.8 %) |
| Coiled-Coil dom. [139-145] | 2 (1.1 %) |
| NLS [146-162] | 9 (4.8 %) |
| Coiled-Coil dom. [163-190] | 14 (7.4 %) |
| Key residues (HCD) | Number of records |
|---|
| C2CH motif [5-5] | 3 (1.6 %) |
| C2CH motif [10-10] | 0 (0 %) |
| DNA binding - [11-11] | 0 (0 %) |
| DNA binding - [24-24] | 3 (1.6 %) |
| DNA binding - [26-26] | 2 (1.1 %) |
| DNA binding - [27-27] | 0 (0 %) |
| DNA binding - [29-29] | 6 (3.2 %) |
| AA interactions [32-32] | 6 (3.2 %) |
| DNA binding - [36-36] | 0 (0 %) |
| DNA binding - [37-37] | 2 (1.1 %) |
| AA interactions [39-39] | 1 (0.5 %) |
| AA interactions [40-40] | 0 (0 %) |
| DNA binding - [42-42] | 0 (0 %) |
| DNA binding - [45-45] | 27 (14.4 %) |
| DNA binding - [48-48] | 0 (0 %) |
| DNA binding - [50-50] | 1 (0.5 %) |
| C2CH motif [54-54] | 3 (1.6 %) |
| C2CH motif [57-57] | 1 (0.5 %) |
| DNA binding - [58-58] | 2 (1.1 %) |
| AA interactions [63-63] | 0 (0 %) |
| AVPTIF motif [76-77] | 2 (1.1 %) |
| DNA binding (AVPTIF) - [78-78] | 0 (0 %) |
| AVPTIF motif [79-81] | 6 (3.2 %) |
| AA interactions [81-81] | 4 (2.1 %) |
| HCF-1 binding [134-137] | 5 (2.7 %) |
| THAP1 dimerization [154-166] | 7 (3.7 %) |
