The UMD-MSH2 mutations database
Structural domains and key residues
This function displays the distribution of small rearrangements in structural domains and key residues of the MSH2 protein. Only mutations/variations found in probands are taken into account.
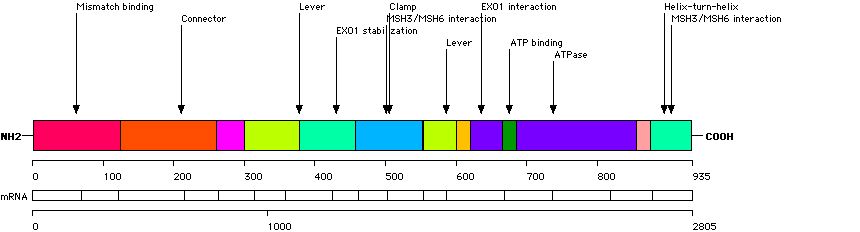
| Structural domain | Number of records |
|---|
| Mismatch binding [1-124] | 731 (22.6 %) |
| Connector [125-297] | 264 (8.2 %) |
| EXO1 stabilization [261-600] | 1361 (42.1 %) |
| Lever [300-456] | 33 (1 %) |
| MSH3/MSH6 interaction [378-625] | 21 (0.6 %) |
| Clamp [457-553] | 16 (0.5 %) |
| Lever [554-619] | 29 (0.9 %) |
| EXO1 interaction [601-671] | 4 (0.1 %) |
| ATPase [620-855] | 383 (11.9 %) |
| ATP binding [666-686] | 175 (5.4 %) |
| Helix-turn-helix [856-934] | 19 (0.6 %) |
| MSH3/MSH6 interaction [875-934] | 122 (3.8 %) |
| Key residues (HCD) | Number of records |
|---|
| [-] | 0 (0 %) |
